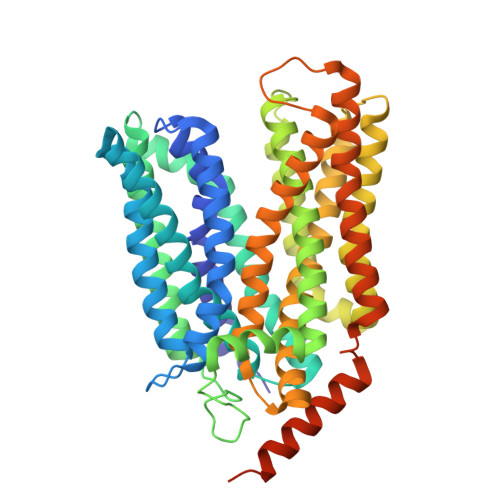
Allosteric effects of the coupling cation in melibiose transporter MelB.
Hariharan, P., Shi, Y., Bakhtiiari, A., Liang, R., Viner, R., Guan, L.(2026) Elife 14
- PubMed: 41604452
- DOI: https://doi.org/10.7554/eLife.108335
- Primary Citation Related Structures:
9OLD, 9OLI, 9OLP, 9OLR - PubMed Abstract:
The major facilitator superfamily (MFS) transporters play significant roles in human health and disease. Salmonella enterica serovar Typhimurium melibiose permease (MelB St ) catalyzes the symport of galactosides with Na + , H + , or Li + and is a prototype of MFS transporters. We published the structures of MelB St in both inward- and outward-facing conformations, bound to galactoside or Na + , and proposed that positive cooperativity of the co-transported solutes is crucial for the symport mechanism. Here, we elucidated the underlying mechanisms by analyzing MelB St dynamics and the effects of melibiose, Na + , or both using hydrogen-deuterium exchange mass spectrometry (HDX-MS). We also refined the determinants of sugar recognition by solving the crystal structures of a uniporter D59C MelB St complexed with melibiose and other sugars, and by identifying a critical water molecule involved in sugar recognition. Our integrated studies, combining structures, HDX-MS, and molecular dynamics simulations, support the conclusion that sugar-binding affinity is directly correlated with protein dynamics. Na + acts as an allosteric activator, reducing the flexibility of dynamic residues in the sugar-binding site and in the cytoplasmic gating salt-bridge network, thereby increasing sugar-binding affinity. This study provides a molecular-level framework of the symport mechanism that could serve as a general model for cation-coupled symporters.
- Department of Cell Physiology and Molecular Biophysics, Center for Membrane Protein Research, School of Medicine, Texas Tech University Health Sciences Center, Lubbock, United States.
Organizational Affiliation: