De novo design of potent CRISPR-Cas13 inhibitors
Taveneau, C., Chai, X.H., D'Silva, J., Bamert, R.S., Hayes, B.K., Calvert, R.W., Curwen, D.J., Munder, F., Martin, L.L., Barr, J.J., Grinter, R., Knott, G.J.To be published.
Experimental Data Snapshot
Starting Models: experimental
View more details
wwPDB Validation 3D Report Full Report
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| AIcrVIA1 | A [auth B] | 83 | synthetic construct | Mutation(s): 0 | 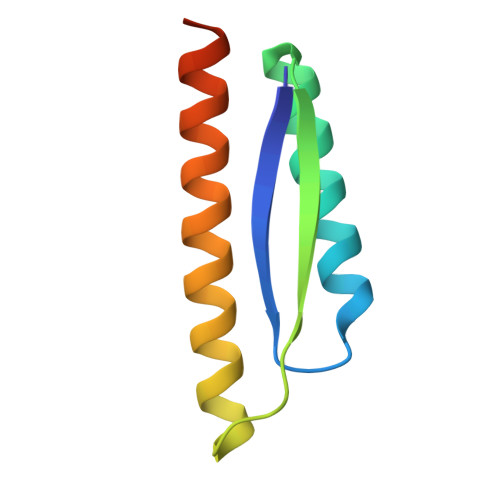 |
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
Sequence AnnotationsExpand | |||||
| |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| CRISPR-associated endoribonuclease Cas13a | B [auth A] | 1,162 | Leptotrichia buccalis | Mutation(s): 0 Gene Names: cas13a, c2c2, Lebu_1799 EC: 3.1 | 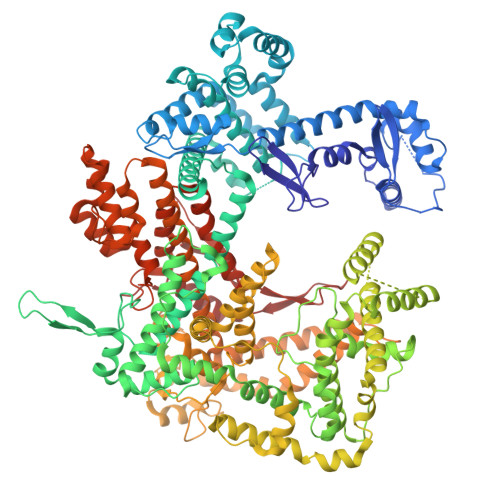 |
UniProt | |||||
Find proteins for C7NBY4 (Leptotrichia buccalis (strain ATCC 14201 / DSM 1135 / JCM 12969 / NCTC 10249 / C-1013-b)) Explore C7NBY4 Go to UniProtKB: C7NBY4 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | C7NBY4 | ||||
Sequence AnnotationsExpand | |||||
| |||||
Entity ID: 3 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Length | Organism | Image | |
| crRNA | 56 | Leptotrichia buccalis | 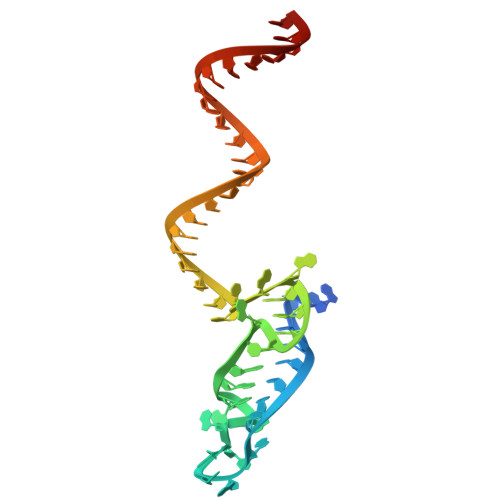 | ||
Sequence AnnotationsExpand | |||||
| |||||
Entity ID: 4 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Length | Organism | Image | |
| Guide complementary activator RNA (aRNA) | 26 | synthetic construct |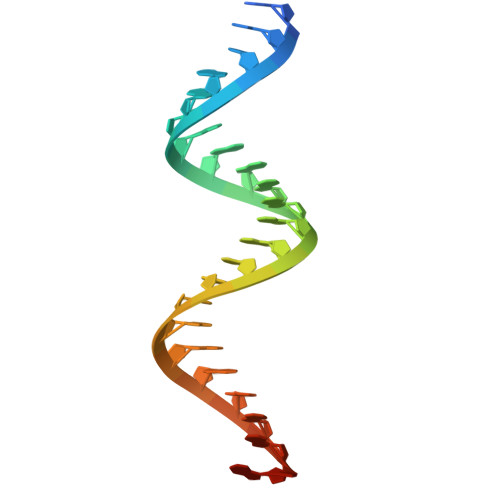  | ||
Sequence AnnotationsExpand | |||||
| |||||
| Task | Software Package | Version |
|---|---|---|
| RECONSTRUCTION | Coot | 0.9.4 |
| RECONSTRUCTION | PHENIX | 1.18.2 |
| MODEL REFINEMENT | PHENIX | 1.18.2 |
| Funding Organization | Location | Grant Number |
|---|---|---|
| Other private | Australia | SMRF2021-276 |