Substrate-engaged TOM complex from yeast
Yang, Y.Q., Wang, G.P.To be published.
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Mitochondrial import receptor subunit TOM40 | A, F [auth I] | 387 | Saccharomyces cerevisiae S288C | Mutation(s): 0 Gene Names: TOM40, ISP42, MOM38, YMR203W, YM8325.04 Membrane Entity: Yes |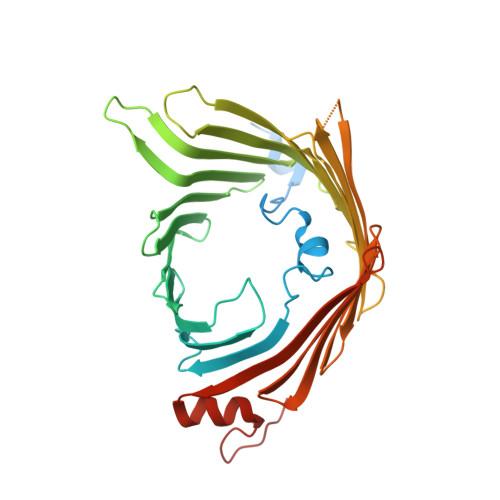  |
UniProt | |||||
Find proteins for P23644 (Saccharomyces cerevisiae (strain ATCC 204508 / S288c)) Explore P23644 Go to UniProtKB: P23644 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P23644 | ||||
Sequence AnnotationsExpand | |||||
| |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Mitochondrial import receptor subunit TOM22 | B, G [auth J] | 152 | Saccharomyces cerevisiae S288C | Mutation(s): 0 Gene Names: TOM22, MAS17, MAS22, MOM22, YNL131W, N1217, N1862 Membrane Entity: Yes |  |
UniProt | |||||
Find proteins for P49334 (Saccharomyces cerevisiae (strain ATCC 204508 / S288c)) Explore P49334 Go to UniProtKB: P49334 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P49334 | ||||
Sequence AnnotationsExpand | |||||
| |||||
Entity ID: 3 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Mitochondrial import receptor subunit TOM5 | C, H [auth K] | 50 | Saccharomyces cerevisiae S288C | Mutation(s): 0 Gene Names: TOM5, MOM8A, YPR133W-A Membrane Entity: Yes |  |
UniProt | |||||
Find proteins for P80967 (Saccharomyces cerevisiae (strain ATCC 204508 / S288c)) Explore P80967 Go to UniProtKB: P80967 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P80967 | ||||
Sequence AnnotationsExpand | |||||
| |||||
Entity ID: 4 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Mitochondrial import receptor subunit TOM6 | D, I [auth L] | 61 | Saccharomyces cerevisiae S288C | Mutation(s): 0 Gene Names: TOM6, ISP6, YOR045W Membrane Entity: Yes |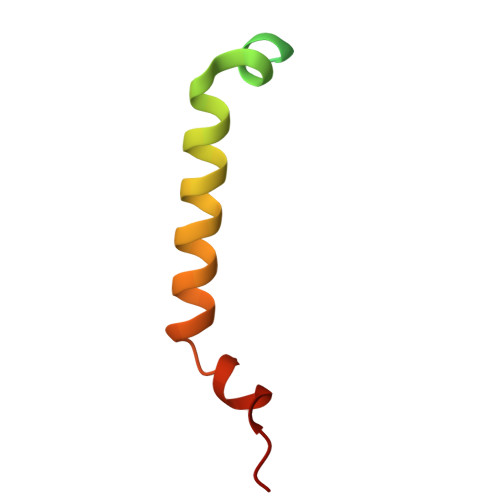  |
UniProt | |||||
Find proteins for P33448 (Saccharomyces cerevisiae (strain ATCC 204508 / S288c)) Explore P33448 Go to UniProtKB: P33448 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P33448 | ||||
Sequence AnnotationsExpand | |||||
| |||||
Entity ID: 5 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Mitochondrial import receptor subunit TOM7 | E, J [auth M] | 60 | Saccharomyces cerevisiae S288C | Mutation(s): 0 Gene Names: TOM7, MOM7, YNL070W, N2378 Membrane Entity: Yes |  |
UniProt | |||||
Find proteins for P53507 (Saccharomyces cerevisiae (strain ATCC 204508 / S288c)) Explore P53507 Go to UniProtKB: P53507 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P53507 | ||||
Sequence AnnotationsExpand | |||||
| |||||
| Ligands 2 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| PC1 Query on PC1 | AB [auth L] CA [auth D] DA [auth D] EA [auth D] FB [auth M] | 1,2-DIACYL-SN-GLYCERO-3-PHOSPHOCHOLINE C44 H88 N O8 P NRJAVPSFFCBXDT-HUESYALOSA-N |  | ||
| UND (Subject of Investigation/LOI) Query on UND | AA [auth C] BA [auth D] BB [auth L] CB [auth L] DB [auth L] | UNDECANE C11 H24 RSJKGSCJYJTIGS-UHFFFAOYSA-N |  | ||
| Task | Software Package | Version |
|---|---|---|
| RECONSTRUCTION | RELION |
| Funding Organization | Location | Grant Number |
|---|---|---|
| National Natural Science Foundation of China (NSFC) | China | 32271269 |