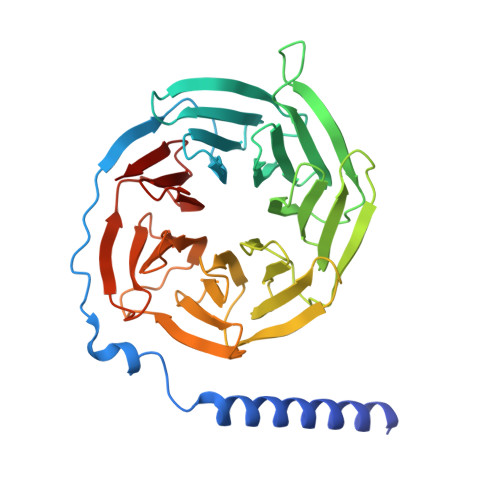
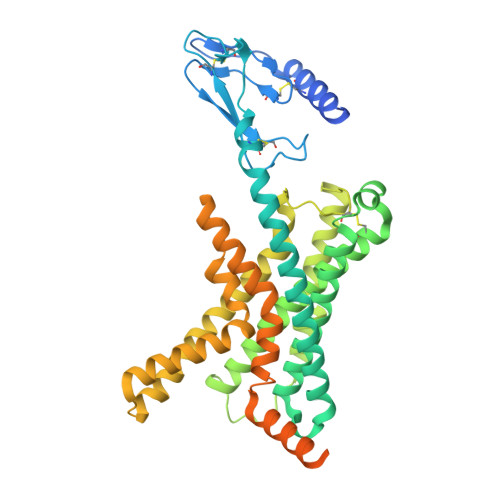
Structural and dynamic features of cagrilintide binding to calcitonin and amylin receptors.
Cao, J., Belousoff, M.J., Johnson, R.M., Keov, P., Mariam, Z., Deganutti, G., Christopoulos, G., Hick, C.A., Reedtz-Runge, S., Glendorf, T., Ballarin-Gonzalez, B., Raun, K., Bayly-Jones, C., Wootten, D., Sexton, P.M.(2025) Nat Commun 16: 3389-3389
- PubMed: 40204768
- DOI: https://doi.org/10.1038/s41467-025-58680-y
- Primary Citation of Related Structures:
9BLB, 9BLC, 9BLW, 9BP3, 9BQ3, 9BTW, 9BUB, 9BUC, 9BUD, 9BUE - PubMed Abstract:
Obesity is a major and increasingly prevalent chronic metabolic disease with numerous comorbidities. While recent incretin-based therapies have provided pharmaceutical inroads into treatment of obesity, there remains an ongoing need for additional medicines with distinct modes of action as independent or complementary therapeutics. Among the most promising candidates, supported by phase 1 and 2 clinical trials, is cagrilintide, a long-acting amylin and calcitonin receptor agonist. As such, understanding how cagrilintide functionally engages target receptors is critical for future development of this target class. Here, we determine structures of cagrilintide bound to Gs-coupled, active, amylin receptors (AMY 1 R, AMY 2 R, AMY 3 R) and calcitonin receptor (CTR) and compare cagrilintide interactions and the dynamics of receptor complexes with previously reported structures of receptors bound to rat amylin, salmon calcitonin or recently developed amylin-based peptides. These data reveal that cagrilintide has an amylin-like binding mode but, compared to other peptides, induces distinct conformational dynamics at calcitonin-family receptors that could contribute to its clinical efficacy.
- Drug Discovery Biology Theme, Monash Institute of Pharmaceutical Sciences, Monash University, Parkville, VIC, Australia.
Organizational Affiliation: