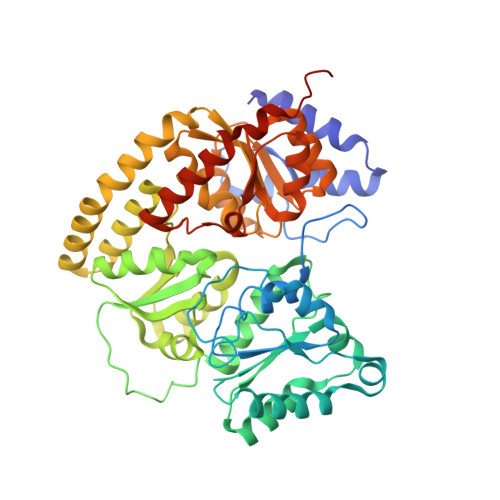
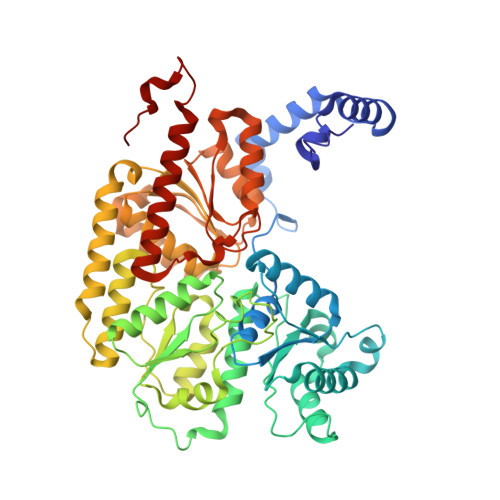
Structures of the nitrogenase complex prepared under catalytic turnover conditions.
Rutledge, H.L., Cook, B.D., Nguyen, H.P.M., Herzik Jr., M.A., Tezcan, F.A.(2022) Science 377: 865-869
- PubMed: 35901182 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/science.abq7641
- Primary Citation Related Structures:
7UT6, 7UT7, 7UT8, 7UT9, 7UTA, 8DPN - PubMed Abstract:
The enzyme nitrogenase couples adenosine triphosphate (ATP) hydrolysis to the multielectron reduction of atmospheric dinitrogen into ammonia. Despite extensive research, the mechanistic details of ATP-dependent energy transduction and dinitrogen reduction by nitrogenase are not well understood, requiring new strategies to monitor its structural dynamics during catalytic action. Here, we report cryo-electron microscopy structures of the nitrogenase complex prepared under enzymatic turnover conditions. We observe that asymmetry governs all aspects of the nitrogenase mechanism, including ATP hydrolysis, protein-protein interactions, and catalysis. Conformational changes near the catalytic iron-molybdenum cofactor are correlated with the nucleotide-hydrolysis state of the enzyme.
- Department of Chemistry and Biochemistry, University of California, San Diego, La Jolla, CA 92093, USA.
Organizational Affiliation: