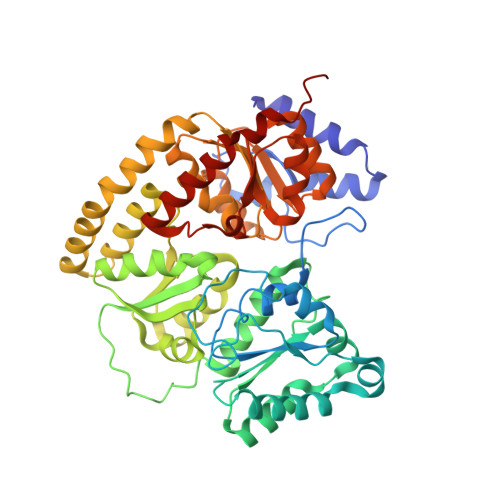
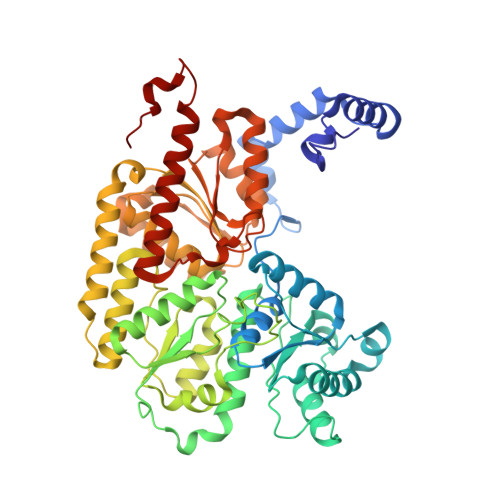
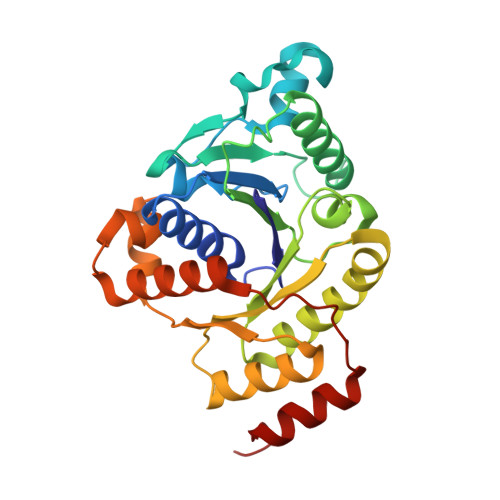
Structural evidence for asymmetrical nucleotide interactions in nitrogenase.
Tezcan, F.A., Kaiser, J.T., Howard, J.B., Rees, D.C.(2015) J Am Chem Soc 137: 146-149
- PubMed: 25522159 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/ja511945e
- Primary Citation Related Structures:
4WZA - PubMed Abstract:
The roles of ATP hydrolysis in electron-transfer (ET) reactions of the nitrogenase catalytic cycle remain obscure. Here, we present a new structure of a nitrogenase complex crystallized with MgADP and MgAMPPCP, an ATP analogue. In this structure the two nucleotides are bound asymmetrically by the Fe-protein subunits connected to the two different MoFe-protein subunits. This binding mode suggests that ATP hydrolysis and phosphate release may proceed by a stepwise mechanism. Through the associated Fe-protein conformational changes, a stepwise mechanism is anticipated to prolong the lifetime of the Fe-protein-MoFe-protein complex and, in turn, could orchestrate the sequence of intracomplex ET required for substrate reduction.
- Department of Chemistry and Biochemistry, University of California, San Diego , La Jolla, California 92093-0356, United States.
Organizational Affiliation: